Our Project
A growing body of data indicates that the propagation of pathogenic protein aggregates across neural systems could be mediated by misfolded protein seeds that are released and taken up by anatomically connected neurons causing disruption of their function. Therefore, blocking this process may help arrest the progression of PD and AD.
This requires a pre-clinical stage of development that has yet to be met. First, it requires in-depth understanding of the biophysical nature of disease-relevant misfolded assemblies to enable the development of tractable molecular and cell-based readouts of their seeding, trafficking and clearance that closely resemble key pathogenic steps in the human condition. On the other end, there is a need to study early oligomerisation events and develop assays of oligomer trafficking inside cells.
The combination of such approaches could form the basis of facile, powerful platforms to map in an unbiased fashion key steps in the propagation cycle or toxicity. An important next step in the trajectory to the development of targeted therapeutics is the availability of tools that allow rapid validation of targets in cellular systems that more closely mimic the interconnected networks of the brain.
Currently this is limited to animal models, which develop pathology over several months. This bottleneck could be circumvented by organotypic cultures and iPSC-derived models with fluorescent tags to enable live imaging of propagation between neurons.
Another consideration is the species barrier, which though not substantial, it increases incubation periods and costs and may impact on the extent of neurodegeneration. This can be addressed by the development of humanized models as well as a combination of minimally invasive and/or peripherally accessible readouts that facilitate the more effective use of animals.
This requires a pre-clinical stage of development that has yet to be met. First, it requires in-depth understanding of the biophysical nature of disease-relevant misfolded assemblies to enable the development of tractable molecular and cell-based readouts of their seeding, trafficking and clearance that closely resemble key pathogenic steps in the human condition. On the other end, there is a need to study early oligomerisation events and develop assays of oligomer trafficking inside cells.
The combination of such approaches could form the basis of facile, powerful platforms to map in an unbiased fashion key steps in the propagation cycle or toxicity. An important next step in the trajectory to the development of targeted therapeutics is the availability of tools that allow rapid validation of targets in cellular systems that more closely mimic the interconnected networks of the brain.
Currently this is limited to animal models, which develop pathology over several months. This bottleneck could be circumvented by organotypic cultures and iPSC-derived models with fluorescent tags to enable live imaging of propagation between neurons.
Another consideration is the species barrier, which though not substantial, it increases incubation periods and costs and may impact on the extent of neurodegeneration. This can be addressed by the development of humanized models as well as a combination of minimally invasive and/or peripherally accessible readouts that facilitate the more effective use of animals.
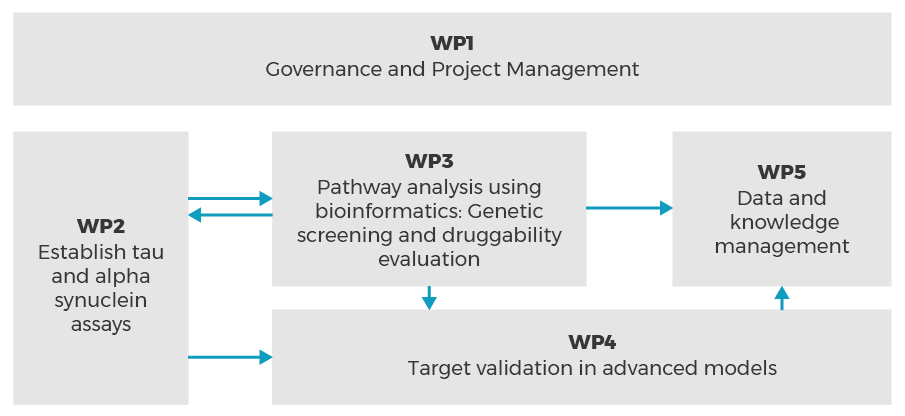
The main scientific part of IMPRiND is structured by a forward moving value chain where tools, molecular assemblies and assays are developed and defined in WP2 which is feeding into WP3 where the target identification and initial target characterization and evaluation of druggability is carried out. The more advanced characterization and target validation is carried out in WP4. Although in general this architecture is outlined as sequential, many of these activities are inter-related, overlapping and may run in parallel.
This project receives funding from the Innovative Medicines Initiative 2 Joint Undertaking (www.imi.europa.eu) under grant agreement No 116060. This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA.
This work is supported by the Swiss State Secretariat for Education‚ Research and Innovation (SERI) under contract number 17.00038.
The opinions expressed and arguments employed herein do not necessarily reflect the official views of these funding bodies.



